Chromatin is defined as a complex of RNA, DNA, and Protein observed in eukaryotic cells. Its prime function lies in the packaging of DNA molecules in a very long denser compact shape withholds the strands from becoming tangled and plays a role in strengthening the DNA during cell division.
In the process of mitosis and meiosis, chromatin helps to separate chromosomes during anaphase. where the typical shapes are observed. The word chromosome has been taken from the Greek word chroma, color, and soma, due to their property of being stained strongly by particular dyes. Likewise, the lightly stained less condense a portion of chromatin termed euchromatin while the darkly stained and highly condensed region of chromatin is termed heterochromatin.
Interesting Science Videos
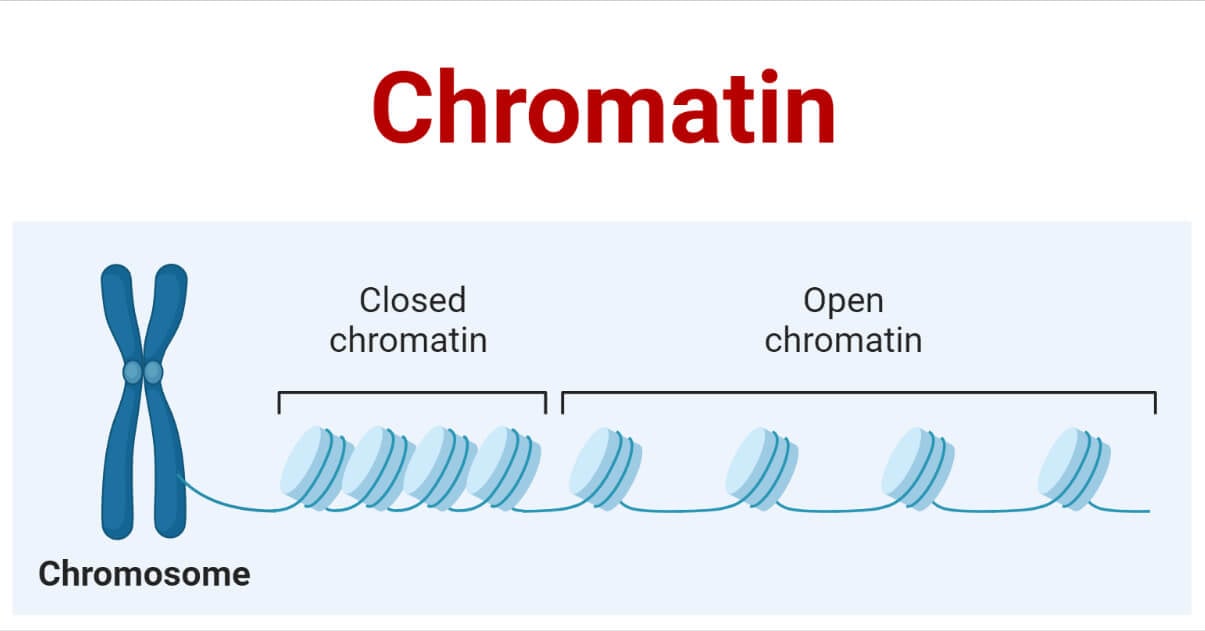
Chromatin Organization
- DNA wraps around the histone proteins making beads on a string structure.
- Several histones wrap into 30nm fiber-like arrays in most of the solid form(heterochromatin)
- A higher level of DNA supercoiling of the 30nm fiber creates the stage of metaphase chromosome. Nucleosome arrays form more complex 30nM high ordered fibers:
The concept of binding of H1 stabilizes higher-order chromatin structure. However, practically, the addition of high salt concentrations of histone H1 results in the formation of 30nm fibers of nucleosomal DNA. Furthermore, in vivo observation represents the other level of DNA compaction.
Consequently, DNA incorporation into the fiber makes the DNA less accessible to other DNA-dependent enzymes like RNA polymerases.
Packaging of DNA into Chromosomes
Model 1: In the solenoid model, the nucleosome DNA forms a superhelix of 6 nucleosomes per turn, where which is supported by biophysical techniques such as electron microscopy and X-ray diffraction. The studies revealed that 30nm fiber has a helical path of ~11nm. The flat surfaces on either side of the histone octamer disc are adjacent to each other, while the DNA surface of the nucleosomes occupies the accessible surface of the superhelix. The linked DNA is found moto be buried inside the mid-region of the superhelix, but it never passes through the axis of the fiber. In another way, the linker DNA wraps around the central axis as the DNA moves from one nucleosome to the next.
Model 2– zig-zag, which is based on the pattern of nucleosomes formed upon H1, are passed through the central axis of the fiber, and the sides and even entry and exit points tend to be more accessible.
Nuclear Scaffold
This is represented by two classes of proteins Topoisomerase 2, is abundantly present in scaffold preparations as well as mitotic chromosomes in purified forms. However, Topo 2 holds the DNA from the base of the loop to make sure that they are isolated from one another topologically.
The SMC class of proteins represents the abundant components of the nuclear scaffold, These proteins play a key component in the machinery that contributes and condenses and holds the sister chromatids after chromosomal duplication. Besides, the association of these proteins serves to enhance the function of the interactions with chromosomal DNA.
The structure of the chromosome lies in the stage of the cell cycle. During the interphase stage, the chromatin loses its permit to access DNA and RNA polymerases which copy and replicate the DNA. DNA consists of genes that are not compacted tightly while closely related with RNA polymerases in a structure called euchromatin while the inactive regions which look more condensed are linked to structural proteins in heterochromatin. Epigenetic alterations include modifications like acetylation, methylation, sumoylation, and sulfation. The chromatin structure and its systems remain a hot cake in the research field.
Eukaryotic Chromosome in Cell Division
Chromatin material is composed of remodeling of nucleosomes, chemical modification of histones or incorporation of variants, non- histones of DNA binding proteins along with noncoding RNAs. However, the affinity is still governed by the post-transcriptional modifications(PTM) by histones for further regulation of transcriptional, recombination, and repair activity. The chromatin structure brings alterations by a distinct class of remodeling enzymes called remodelers of chromatin.
However, the few structural changes throughout the cell cycle undergo changes. Histone proteins are the packers and coordinators of the chromatin and can be altered by numerous post-translational changes. Majorly important modifications occur in the tail region of histone proteins. Along, the consequences in chromatin availability and compaction depend both on the amino acid and the kind of modification involved. Apart from that, acetylation by histone proteins ends up in loosening and rising accessibility for different events like duplication and transcription. The field of tumorigenesis has already marked the proteins as chromatin modifiers and remodelers. The proteins obtained by this fusion would have a profound impact on the transcription machinery, which might explain why soft tissue tumors with undifferentiated, as exemplified by undifferentiated cell sarcoma or display disparate lines of differentiation, such as synovial sarcoma or ossifying fibromyxiod tumor with PHF1 fusions.
Techniques for analysis of chromatin
The two advancements are discussed below briefly:
- Dnase1 Hypersensitivity: This method considers the recognition of the position of monitoring regions, based n the genomics wide sequencing which further cleaves the entire structure by Dnase1. However, DNAse1 occupied a minor groove when incubated with DNA resulting in phosphodiester bonds on both strands by separating the DNA. However, it results in many fragments of DNA making it a more effective nuclease by adding EDTA, or EGTA.
- DNA Footprinting: This method evaluates the identification of DNA- binding proteins in certain regions. An inhibitory detection assay is performed by PAGE electrophoresis, if a specific protein, ligand, or any of the transcriptional factors bind to the DNA or not. Certain types of DNA modifiers like free radicals would cut the DNA, ligand linked DNA describes a differential banding pattern in a gel form or as a fingerprint or as a certain gap.
The following methods include
a) amplification of the gene of interest
b) Radio or Fluorolabelled DNA
c) Protein with DNA incubation or commonly called Immunoprecipitation
d) Modification of DNA by DNA modifiers
e) DNA fragments by gel electrophoresis
f) Interpretation of results - Chromatin- Immunoprecipitation Sequencing: The process used to analyze the interaction of proteins with DNA. Likewise, the binding sites of DNA along with the proteins were identified by the chromatin immunoprecipitation along with parallel DNA sequencing.
- Formaldehyde–assisted isolation of certain regulatory elements sequencing
- Assay for transposable accessible chromatin sequencing
- Micrococcal nuclease sequencing: This method aids in the depth study of nucleosomes by digesting chromatins. However, this depicts the position of nucleosomes throughout the genome.
Microscopic Observation
Upon staining, eukaryotic nuclear DNA is observed in nondividing cells in two different states, heterochromatin( dark areas) can be observed along the inner side of the nuclear envelope as well as inside the nucleus, and euchromatin ( light areas) throughout the nucleus. While speaking of heterochromatin, it contains transcriptionally inactive genes which protect it from tightly packed nucleases. One of the X chromosomes is silenced while the other remains active in euchromatin, particularly in humans.
Euchromatin occupies 90% of the genome which looks unfolded resembling beads on the string found only in bacteria. These are active transcriptionally upon binding of RNA polymerase. In addition to eukaryotic cells, DNA is organized as a long linear structure called chromosomes with complex bound proteins. The proteins bind to and condense the DNA molecule preventing it from tangling. Onset before the cell division, replication brings about an identical set of chromosomes called sister chromatids binding at the center of each to a common structure called the centromere. Normally human cell contains 23 pairs of chromosomes, out of which 22 pairs are autosomes while one pair is a sex chromosome which differs in males and females. Each species comprises a unique chromosomal complement. However, the total length of the human genome is over 3 billion base pairs including mitochondrial DNA. Eukaryotic species(protists, fungi, plants, and animals) store the content inside a cell nucleus as linear nuclear DNA. Likewise, mitochondria and chloroplast comprise circular DNA( no ends). Lower organisms such as archaea and bacteria do not have organelle structures and thus, store their DNA in a region of cytoplasm called as nucleoid region. Although prokaryotic chromosomes consist of double-stranded circular DNA.
In a recent finding, a transcription mediated by a glucocorticoid receptor(GR) was found to play a pivot role in the treatment of diseases. The data by Wang et al, 2021 described that the binding of glucocorticoid activates regulatory elements in podocytes which seem like a primary medication for podocytopathy. The interaction of GR at the transcription level was found to be indispensable to the effective functioning of GC in podocytes. Conclusively, this study has demonstrated the need for considering transcriptional activators in order to perfectly map regulatory networks of GR.
Advantages
- It exactly locates the binding region of the particular ligand
- It is affordable commercially.
- This technique is powerful enough to segregate many fragments.
Disadvantages
- This is the most conventional, time-consuming, and laborious.
- The banding resolution seems very low
- The use of radioactivity is more hazardous.
Conclusion
Despite the crucial role in many cellular and molecular activities of DNA, the defective functioning of chromatin remodeling leads to a broad spectrum of many human disorders like cancer, birth defects, and developmental disorders. The reason behind this lies in the complexity of DNA replication, recombination, repair, and growth with which many genes reprogramming in case of cancer progression. The structural study could be one of the keys to opening the precise molecular depth along with epigenetics in human diseases. This would pave the way to the possible therapeutics of proteins involved in the nearby future.
References
- Cote J, Quinn J, Workman JL, Peterson CL. Stimulation of GAL4 derivative binding to nucleosomal DNA by the yeast SWI/SNF complex. Science 1994; 265:53-60; PMID:8016655; http://dx.doi.org/10.1126/science.8016655.
- Wang, H., Duan, A., Zhang, J. et al. Glucocorticoid receptor wields chromatin interactions to tune transcription for cytoskeleton stabilization in podocytes. Commun Biol4, 675 (2021). https://doi.org/10.1038/s42003-021-02209-8
