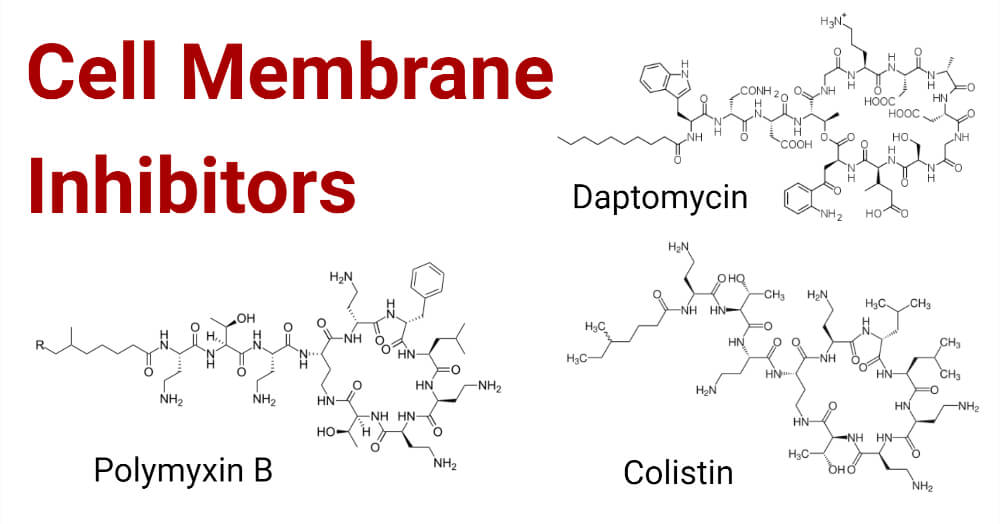
In this era, bacterial membrane became the new target for the antimicrobial agent because anionic lipids are present in bacterial surfaces and the antimicrobials are cationic so they show greater reactivity towards membranes. Gram-positive and Gram-negative bacteria are morphologically and functionally different from one another. Gram-negative bacteria contain two membranes, the outer membrane and, cytoplasmic membrane, which are made up of lipid components. The major lipid components present in the cell membrane are phosphatidylglycerol, phosphatidylethanolamine and, cardiolipin. Different bacterial species have a different proportion of these lipids which act differently in their way. Different antimicrobials targets cell membrane for bacterial lysis which includes Polymyxins of the Polypeptide class and Daptomycin of Lipopeptide which is a newer class of antimicrobial agent.
- The cell membrane has selective permeability which plays important role in the ATP cycle of microorganisms.
- Cell membranes are made up of fatty acids which are synthesized in cells
- Polymyxins, the first line of the drug consists of fifteen different variations. Among other polymyxins, polymyxin E is also known as Colistin isolated from Paenibacillus polymyxa in 1947 and polymyxin M also called mattacin and polymyxin B are mostly used.
- Polymyxin B and Polymyxin B have quite similar structures which differ by only one amino acid.
- Polymyxins have bactericidal properties.
- Polymyxins are produced from the non-ribosomal peptide synthetase system.
- Daptomycin is cyclic polypeptides that are produced naturally belonging to lipopeptides which consist of peptide core with lipid tail in its structure.
- Daptomycin was produced by Streptomyces roseosporus with is used against Gram-positive organisms.
- In 1953, the first daptomycin are isolated after the discovery of amphomycin. It has a unique structure that consists of a 13 member amino acid cyclic chain with a decanoyl side chain.
- Both polymyxins and Daptomycins cause depolarization and potassium ion efflux resulting in bacterial cell death.
- Daptomycin is used to treat infections caused by both Gram-positive and Gram-negative bacteria including Vancomycin-resistant Staphylococcus aureus. It disrupts membrane potential by depolarization where potassium ions are released causing cell lysis.
Mechanism of action of Cell Membrane Inhibitors
Mainly antimicrobials target phospholipids in the cell membrane Antimicrobials alters the physical property which includes intrinsic curvature or fluidity within the cell. Clustering of anionic lipids with cationic agents can be lethal to bacteria causing cell lysis.
- Daptomycin: Daptomycin shows a bactericidal property. It alters the homeostasis by reacting with the phospholipids of the cell membrane. After insertion of daptomycin into the cell membrane, the daptomycin calcium complex gets oligomerized in the outer leaf-like structure. Oligomerization of daptomycin calcium complex aids in bactericidal activity. As daptomycin’s are cationic antimicrobial agents contain pores that have high permeability to sodium, potassium, and other metal ions. Cardiolipin aids in the translocation of daptomycin from the outer structure to the inner structure. Cardiolipin plays important role in homeostasis. Hence, enrichment of liposomes helps to prevent translocation disrupting the formation of pores might prevent the antibacterial activity of daptomycin. Daptomycin also shows little effect in DNA, RNA, and protein synthesis. Followed by depolarization and potassium ion efflux. Leakage of ions and loss of cell membrane potential is the result of bacterial homeostasis that causes cellular damage and death.
- Polymyxins: Permeabilization or disruption of a cell membrane is considered the main mode of action of this category of antibiotics. Polymyxins target the outer membrane of Gram-negative organisms. The outer membrane is more permeable as compared to the cell membrane as it contains pores. Diaminobutyric acid residues, divalent ions, and lipid A are shifted from phosphate groups of lipids which result in the destabilization of lipopolysaccharide. This results in the increases in permeability of membrane causing leakage of cytoplasmic content and causes cell death. Another mechanism involved is the endotoxin effect. Polymyxins can bind and neutralize lipopolysaccharide. And another mode of action includes inhibition of viral respiratory enzymes. Disruption in the cytoplasmic membrane affects cell division and synthesis.
Examples of Cell Membrane Inhibitors
- Colistin
- Daptomycin
Mechanism of resistance of Cell Membrane Inhibitors
- As polymyxins are used to treat multidrug-resistant infections, resistivity to these antimicrobials is of great concern. An Increase in drug efflux, mutation, alteration of the porin pathway is some of the reasons causing resistance. Lipopolysaccharide modification in Gram-negative organisms is governed by two-component PhoP /Q and PmrA/B regulatory systems. These regulatory systems allow the expression of a gene which code for magnesium transport that modifies LPS. Different genes which are encoded chromosomally are responsible for producing LPS are responsible for showing resistance towards Polymyxin B. The most common cause for the resistance is the modification of LPS through cationic substitution. Plasmid-mediated resistance in polymyxins is caused by mcr genes.
- Resistance to daptomycin is also of great concern. Membrane homeostasis, alteration of regulatory systems are associated with the resistivity of organisms towards daptomycin. Genes which cause daptomycin resistant are gene that is involved in phospholipid mechanisms.
The enzymes that join peptidoglycan’s two molecules to make cardiolipin are responsible for cardiolipin synthesis and cardiolipin synthase. Thus, it is important to know that mutation producing changes in enzyme plays important role in daptomycin resistant by altering the ratio of peptidoglycan to cardiolipin in the cell membrane. Membrane fluidity is also an important factor that influences the daptomycin resistance in staphylococci.
References
- Activity PA (2017). crossm. 30(2), 557–596.
- Epand R M, Walker C, Epand, RF, Magarvey N A and Hilpert K (2016). Biochimica et Biophysica Acta Molecular mechanisms of membrane targeting antibiotics . BBA – Biomembranes, 1858(5), 980–987. https://doi.org/10.1016/j.bbamem.2015.10.018
- From, Introductory Chapter: The Action Mechanisms of Antibiotics and Antibiotic Resistance | IntechOpen.
- Kola M, Trimble, M. J and Mlyna P (2016). Polymyxin : Alternative Mechanisms of Action. 1–22.
- Miller W R, Bayer A S and Arias C A (2016). Mechanism of Action and Resistance to Daptomycin in Staphylococcus aureus and Enterococci. 1–16.
- Steenbergen J N, Alder J, Thorne G M and Tally FP (2005). Daptomycin: a lipopeptide antibiotic for the treatment of serious Gram-positive infections. 55(3):283–288. https://doi.org/10.1093/jac/dkh546
- Tran TT, Munita J M, Arias C. A., Santiago A De, Genetics M and Unit A R. (2016). HHS Public Access. (1): 32–53. https://doi.org/10.1111/nyas.12948.Mechanisms
- Yu Z, Qin W, Lin J, Fang S and Qiu J (2015). Antibacterial Mechanisms of Polymyxin and Bacterial Resistance. 2015.
